UPDATE: See Carl Boettiger’s functions/package at Github for searching Treebase here.
Treebase is a great resource for phylogenetic trees, and has a nice interface for searching for certain types of trees. However, if you want to simply download a lot of trees for analyses (like that in Davies et al.), then you want to be able to access trees in bulk (I believe Treebase folks are working on an API though). I wrote some simple code for extracting trees from Treebase.org.
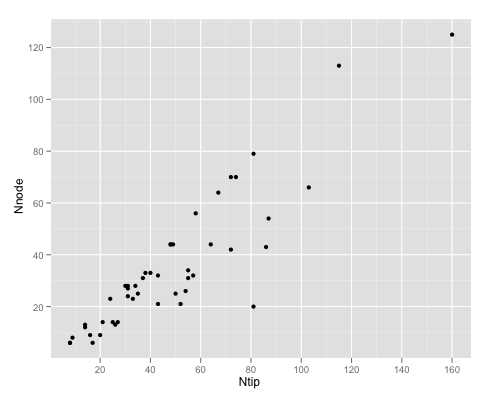
It reads an xml file of (in this case consensus) URL’s for each tree, parses the xml, makes a vector of URL’s, reads the nexus files with error checking, remove trees that gave errors, then a simple plot looking at metrics of the trees.
Is there an easier way to do this?
(note: code is gone, was at https://gist.github.com/953468.js?file=treebase_code.R)